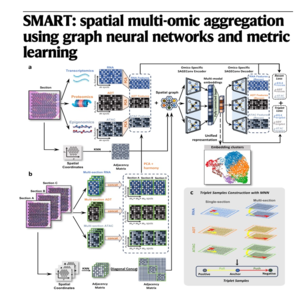
The paper introduces SMART, an unsupervised deep learning framework designed to integrate spatial multi-omics data into a unified representation. By combining graph neural networks with metric learning, the model effectively captures both local spatial contexts and long-distance biological similarities across various omics layers. The researchers developed a variant called SMART-MS to align data across multiple tissue sections while correcting for batch effects. Benchmarking demonstrates that SMART offers superior computational efficiency, scalability, and accuracy in identifying anatomical structures compared to existing methods. Ultimately, this tool provides a versatile solution for analyzing complex tissue microenvironments at various spatial resolutions.
References:
- Du Z, Chen Q, Huang W, et al. SMART: spatial multi-omic aggregation using graph neural networks and metric learning[J]. Nature Communications, 2026, 17(1): 2876.


 1
1 0
0