This study introduces a comprehensive placental transcriptome reference developed through long-read RNA sequencing, uncovering a level of transcriptional complexity previously unrecognized in this organ. By identifying over 37,000 high-confidence isoforms, including thousands of previously unannotated transcripts, the researchers challenge the notion that the placenta is a genomic void. The authors demonstrate that this tissue-specific reference significantly reduces quantification uncertainty in existing datasets compared to generic adult-tissue annotations. Utilizing this refined resolution, the study reveals that specific placental isoforms—particularly of the CSH1 gene—mechanistically mediate the effects of gestational diabetes on newborn birth weight. These findings underscore the critical importance of isoform-level analysis for understanding maternal-fetal health and the developmental origins of disease. Overall, the resource provides a powerful framework for re-evaluating how environmental exposures and genetic factors shape pregnancy outcomes.
References:
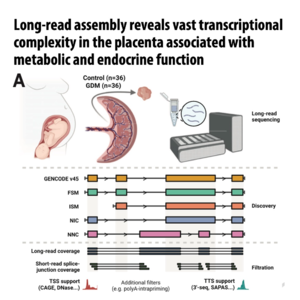
- Bresnahan S T, Yong H E J, Nemani A, et al. Long-read assembly reveals vast transcriptional complexity in the placenta associated with metabolic and endocrine function[J]. Nature Communications, 2026.


 1
1 0
0